It is out: Feralisation targets different genomic loci to domestication in the chicken. This is the second of our papers on the Kauai feral and admixed chicken population, and came out a few days ago.
The Kauai chicken population is kind of famous: you can find them for instance on Flickr, or on YouTube. We’ve previously looked at their plumage, listened to the roosters’ crowings, and sequenced mitochondrial DNA to investigate their origins. Based on this, we concur with the common view that the chickens of Kauai probably are a mixture of feral birds of domestic origin and wild Junglefowl. The Kauai chickens look and sound like a mix of wild and domestic, and we found mitochondrial DNA of two haplogroups, one of which (called D) is typical in ancient chicken DNA from Pacific islands (Gering et al 2015).
In this paper, we looked at the rest of the genome of the same chickens — you didn’t think we sequenced the whole thing just to look at the mitochondrion plus a subset of markers, did you? We turn to population genomics, and a family of methods called selective sweep mapping, to search for regions of their genome that show signs of being affected by natural selection. This lets us: 1) draw pretty rainbow plots such as this one …
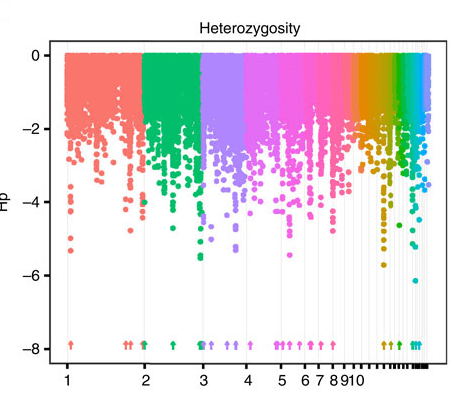
(Figure 1a from the paper in question, Johnsson & al 2016. cc:by The chromosomes have been laid out on the horizontal axis with different colours, and split into windows of 40 kb. Each dot represents the heterozygosity of that windows. For all the details, see the paper.)
… 2) highlight a regions of the genome that may have been selected during feralisation on Kauai (these are the icicles in the graph, highligthed by arrows); 3) conclude that the regions that look like they’ve been selected in feralisation overlap very little with the ones that look like they’ve been selected in chicken domestication. Hence the title.
That was the main result, but of course we also look at what genes are highlighted. Mostly we have no idea how they may contribute to feralisation, but a couple of regions overlap with those that we’ve previously found in genetic mapping of comb size and egg laying in our wild-by-domestic intercross. We also compare the potentially selected regions to domestic chicken sequences.
Last year, Ewen Callaway visited Dominic Wright, Eben Gering and Rie Henriksen on the last fieldtrip to Kauai. The article, When chickens go wild, was published in Nature News in January, and it explains a lot of the ideas nicely. This paper was submitted by then, so the samples they gathered on that trip do not feature in it. But, spoiler alert: there is more to come. (I don’t know what role I personally will play, but that is less important.)
As you may have guessed if you looked at the author list, this was a collaboration between quite a lot of people in Linköping, Michigan, London, and Victoria. Thanks to all involved! This was great fun, and for those of you who like this sort of thing, I hope the paper will be an interesting read.
Literature
M. Johnsson, E. Gering, P. Willis, S. Lopez, L. Van Dorp, G. Hellenthal, R. Henriksen, U. Friberg & D. Wright. (2016) Feralisation targets different genomic loci to domestication in the chicken. Nature Communications. doi:10.1038/ncomms12950












