This is a relatively recent preprint on how correlations between genetic variants can limit the response to selection, with experimental evolution in bacteria.
Experimental evolution and selection experiments live on the gradient from modelling to observations in the wild. Experimental evolution researchers can design the environments and the genotypes to pose problems for evolution, and then watch in real time as organisms solve them. With sequencing, they can also watch how the genome responds to selection.
In this case, the problem posed is how to improve a particular cellular function (”module”). The researcher started out with engineered Escherichia coli that had one component of their translation machinery manipulated: they started out with only one copy of an elongation factor gene (where E.coli normally has two) that could be either from another species, an reconstructed ancestral form, or the regular E.coli gene as a control.
Then, they sequenced samples from replicate populations over time, and looked for potentially adaptive variants: that is, operationally, variants that had large changes in frequency (>20%) and occurred in genes that had more than one candidate adaptive variant.
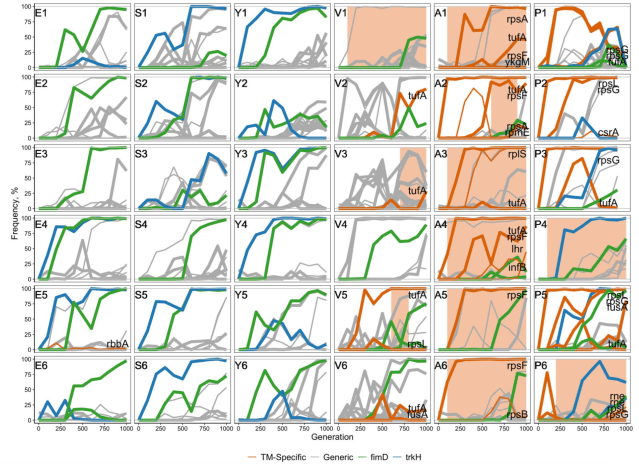
Finally, because they looked at what genes these variants occurred in. Were they related to translation (”TM-specific” as they call it) or not (”generic”). That gave them trajectories of potentially adaptive variants like this. The horizontal axis is time and the vertical frequency of the variant. The letters are populations of different origin, and the numbers replicates thereof. The colour shows the classification of variants. (”fimD” and ”trkH” in the figure are genes in the ”generic” category that are of special interest for other reasons. The orange–brown shading signifies structural variation at the elongation factor gene.)
This figure shows their main observations:
- V, A and P populations had more adaptive variants in translation genes, and also worse fitness at the start of the experiment. This goes together with improving more during the experiment. If a population has poor translation, a variant in a translation gene might help. If it has decent translation efficiency already, there is less scope for improvement, and adaptive variants in other kinds of genes happen more often.
We found that populations whose TMs were initially mildly perturbed (incurring ≲ 3% fitness cost) adapted by acquiring mutations that did not directly affect the TM. Populations whose TM had a moderately severe defect (incurring ~19% fitness cost) discovered TM-specific mutations, but clonal interference often prevented their fixation. Populations whose TMs were initially severely perturbed (incurring ~35% fitness cost) rapidly discovered and fixed TM-specific beneficial mutations.
- Adaptive variants in translation genes tended to increase fast and early during the experiment and often get fixed, suggesting that they have larger effects than. Again, the your translation capability is badly broken, a large-effect variant in a translation gene might help.
Out of the 14 TM-specific mutations that eventually fixed, 12 (86%) did so in the first selective sweep. As a result, an average TM-specific beneficial mutation reached fixation by generation 300 ± 52, and only one (7%) reached fixation after generation 600 … In contrast, the average fixation time of generic mutations in the V, A and P populations was 600 ± 72 generations, and 9 of them (56%) fixed after the first selective sweep
- After one adaptive variant in a translation gene, it seems to stop at that.
The question is: when there aren’t further adaptive variants in translation genes, is that because it’s impossible to improve translation any further, or because of interference from other variants? They use the term ”evolutionary stalling”, kind of asexual linked selection. Because variants occur together, selection acts on the net effect of all the variants in an individual. Adaptation in a certain process (in this case translation) might stall, if there are large-effect adaptive variants in other, potentially completely unrelated processes, that swamp the effect on translation.
They argue for three kinds of indirect evidence that the adaptation in translation has stalled in at least some of the populations:
- Some of the replicate populations of V didn’t fix adaptive translation variants.
- In some populations, there were a second adaptive translation variant, not yet fixed.
- There have been adaptive translation mutations in the Long Term Evolution Experiment, which is based on E.coli with unmanipulated translation machinery.
Stalling depends on large-effect variants, but after they have fixed, adaptation might resume. They use the metaphor of natural selection ”shifting focus”. The two non-translation genes singled out in the above figure might be examples of that:
While we did not observe resumption of adaptive evolution in [translation] during the duration of this experiment, we find evidence for a transition from stalling to adaptation in trkH and fimD genes. Mutations in these two genes appear to be beneficial in all our genetic backgrounds (Figure 4). These mutations are among the earliest to arise and fix in E, S and Y populations where the TM does not adapt … In contrast, mutations in trkH and fimD arise in A and P populations much later, typically following fixations of TM-specific mutations … In other words, natural selection in these populations is initially largely focused on improving the TM, while adaptation in trkH and fimD is stalled. After a TM-specific mutation is fixed, the focus of natural selection shifts away from the TM to other modules, including trkH and fimD.
This is all rather indirect, but interesting. Both ”the focus of natural selection shifting” and ”coupling of modules by the emergent neutrality threshold” are inspiring ways to think about the evolution of genetic architecture, and new to me.
Literature
Venkataram, Sandeep, et al. ”Evolutionary Stalling and a Limit on the Power of Natural Selection to Improve a Cellular Module.” bioRxiv (2019): 850644.