As a wide-eyed PhD student, I read a lot of papers about gene expression networks and was mightily impressed by their power. You can see where this is going, can’t you?
Someone on Twitter talked about their doubts about gene networks: how networks ”must” be how biology works, but that they weren’t sure that network methods actually had helped genetics that much, how there are compelling annotation term enrichments, and individual results that ”make sense”, but not many hard predictions. I promise I’m not trying to gossip about them behind their back, but I couldn’t find the tweets again. If you think about it, however, I don’t think genes must form networks at all, quite the opposite. But there are probably reasons why the network idea is so attractive.
(Edit: Here is the tweet I was talking about by Jeffrey Barrett! Thanks to Guillaume Devailly for pointing me to it.)
First, network representations are handy! There are all kinds of things about genes that can be represented as networks: coexpression, protein interactions, being mentioned in the same PubMed abstract, working on the same substrate, being annotated by the same GO term, being linked in a database such as STRING which tries to combine all kinds of protein–protein interactions understood broadly (Szklarczyk & al 2018), differential coexpression, co-differential expression (Hudson, Reverter & Dalrymple 2009), … There are all kinds of ways of building networks between genes: correlations, mutual information, Bayesian networks, structural equations models … Sometimes one of them will make an interesting biological phenomena stand out and become striking to the eye, or to one of the many ways to cluster nodes and calculate their centrality.
Second, networks are appealing. Birgitte Nerlich has this great blog post–On books, circuits and life–about metaphors for gene editing (the book of life, writing, erasing, cutting and editing) and systems biology (genetic engineering, circuits, wiring, the genetic program). Maybe the view of gene networks fits into the latter category, if we imagine that the extremely dated analogy with cybernetics (Peluffo 2015) has been replaced with the only slightly dated idea of a universal network science. After Internet and Albert, Jeong & Barabási (1999), what could be more apt than understanding genes as forming networks?
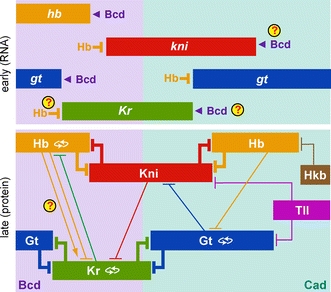
I think it’s fair to say that for genes to form networks, the system needs to be reasonably well described by a graph of nodes and edges. If you look at systems of genes that are really well understood, like the gap gene ”network”, you will see that they do not look like this at all. Look at Fig 3 in Jaeger (2011). Here, there is dynamic and spatial information not captured by the network topology that needs to be overlaid for the network view to make sense.
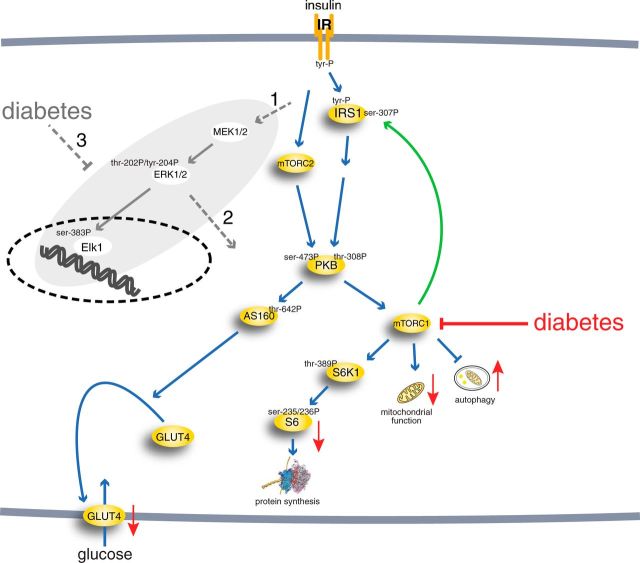
Or look at insulin signalling, in Fig 1 of Nyman et al (2014). Here, there are modified versions of proteins, non-gene products such as glucose and the plasma membrane, and again, dynamics, including both RNA and protein synthesis themselves. There is no justification for assuming that any of that will be captured by any topology or any weighting of genes with edges between them.
We are free to name biological processes networks if we want to; there’s nothing wrong with calling a certain process and group of related genes ”the gap gene network”. And we are free to use any network representation we want when it is useful or visually pleasing, if that’s what we’re going for. However, genes do not actually form networks.
Literature
Szklarczyk, D, et al. (2018) STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic acids research.
Hudson, N. J., Reverter, A., & Dalrymple, B. P. (2009). A differential wiring analysis of expression data correctly identifies the gene containing the causal mutation. PLoS computational biology, 5(5), e1000382.
Peluffo, A. E. (2015). The ”Genetic Program”: behind the genesis of an influential metaphor. Genetics, 200(3), 685-696.
Albert, R., Jeong, H., & Barabási, A. L. (1999). Diameter of the world-wide web. Nature, 401(6749), 130.
Jaeger, J. (2011). The gap gene network. Cellular and Molecular Life Sciences, 68(2), 243-274.
Nyman, E., Rajan, M. R., Fagerholm, S., Brännmark, C., Cedersund, G., & Strålfors, P. (2014). A single mechanism can explain network-wide insulin resistance in adipocytes from obese patients with type 2 diabetes. Journal of Biological Chemistry, 289(48), 33215-33230.