A few weeks ago, I gave a talk at the Edinburgh R users group EdinbR on the RAGE paper. Since this is an R meetup, the talk concentrated on the mechanics of genetic data simulation and with the paper as a case study. I showed off some of what Chris Gaynor’s AlphaSimR can do, and how we built on that to make the specifics of this simulation study. The slides are on the EdinbR Github.
Genetic simulation is useful for all kinds of things. Sure, they’re only as good as the theory that underpins them, but the willingness to try things out in simulations is one of the things I always liked about breeding research.
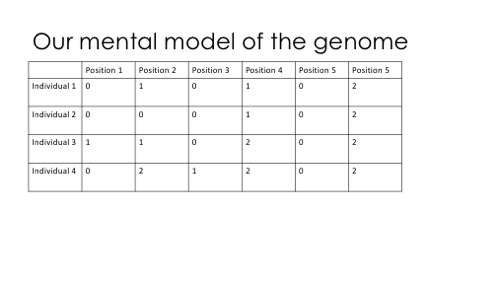
This is my description of the logic of genetic simulation: we think of the genome as a large table of genotypes, drawn from some distribution of allele frequencies.
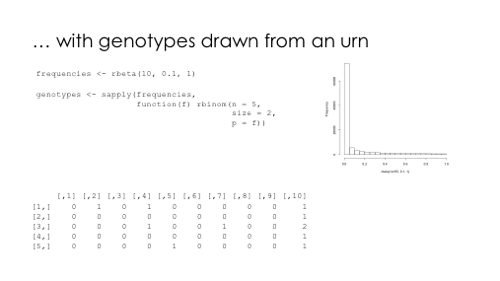
To make an utterly minimal simulation, we could draw allele frequencies from some distribution (like a Beta distribution), and then draw the genotypes from a binomial distribution. Done!
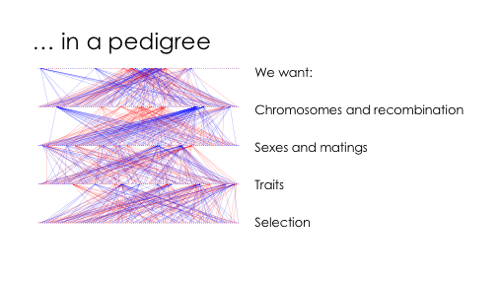
However, there is a ton of nuance we would like to have: chromosomes, linkage between variants, sexes, mating, selection …
AlphaSimR addresses all of this, and allows you to throw individuals and populations around to build pretty complicated designs. Here is the small example simulation I used in the talk.
library(AlphaSimR)
library(ggplot2)
## Generate founder chromsomes
FOUNDERPOP <- runMacs(nInd = 1000,
nChr = 10,
segSites = 5000,
inbred = FALSE,
species = "GENERIC")
## Simulation parameters
SIMPARAM <- SimParam$new(FOUNDERPOP)
SIMPARAM$addTraitA(nQtlPerChr = 100,
mean = 100,
var = 10)
SIMPARAM$addSnpChip(nSnpPerChr = 1000)
SIMPARAM$setGender("yes_sys")
## Founding population
pop <- newPop(FOUNDERPOP,
simParam = SIMPARAM)
pop <- setPheno(pop,
varE = 20,
simParam = SIMPARAM)
## Breeding
print("Breeding")
breeding <- vector(length = 11, mode = "list")
breeding[[1]] <- pop
for (i in 2:11) {
print(i)
sires <- selectInd(pop = breeding[[i - 1]],
gender = "M",
nInd = 25,
trait = 1,
use = "pheno",
simParam = SIMPARAM)
dams <- selectInd(pop = breeding[[i - 1]],
nInd = 500,
gender = "F",
trait = 1,
use = "pheno",
simParam = SIMPARAM)
breeding[[i]] <- randCross2(males = sires,
females = dams,
nCrosses = 500,
nProgeny = 10,
simParam = SIMPARAM)
breeding[[i]] <- setPheno(breeding[[i]],
varE = 20,
simParam = SIMPARAM)
}
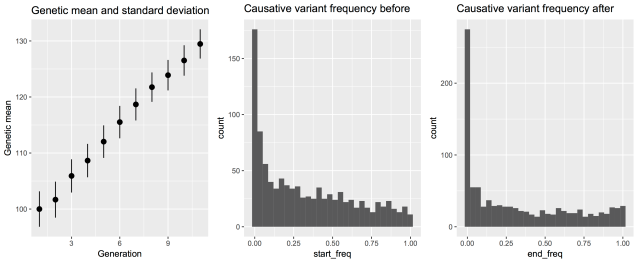
## Look at genetic gain and shift in causative variant allele frequency
mean_g <- unlist(lapply(breeding, meanG))
sd_g <- sqrt(unlist(lapply(breeding, varG)))
plot_gain <- qplot(x = 1:11,
y = mean_g,
ymin = mean_g - sd_g,
ymax = mean_g + sd_g,
geom = "pointrange",
main = "Genetic mean and standard deviation",
xlab = "Generation", ylab = "Genetic mean")
start_geno <- pullQtlGeno(breeding[[1]], simParam = SIMPARAM)
start_freq <- colSums(start_geno)/(2 * nrow(start_geno))
end_geno <- pullQtlGeno(breeding[[11]], simParam = SIMPARAM)
end_freq <- colSums(end_geno)/(2 * nrow(end_geno))
plot_freq_before <- qplot(start_freq, main = "Causative variant frequency before")
plot_freq_after <- qplot(end_freq, main = "Causative variant frequency after")
This code builds a small livestock population, breeds it for ten generations, and looks at the resulting selection response in the form of a shift of the genetic mean, and the changes in the underlying distribution of causative variants. Here are the resulting plots: