This paper (Betts et al. 2021) came out about a month ago investigating whether ecology and evolution papers explicitly state mechanistic hypotheses, and arguing that they ought to, preferably multiple alternative hypotheses. It advocates the particular flavour of hypothetico–deductivism expressed by Platt (1964) as ”strong inference”.
The key idea in Platt’s (1964) account of strong inference, that distinguishes it from garden variety accounts of scientific reasoning, is his emphasis on multiple alternative hypotheses and experiments that distinguish between them. He describes science progressing like a decision tree, with experiments as branching points — a ”conditional inductive tree”. He also emphasises theory construction, as he approvingly quotes biologists on the need to think hard about what possibilities there are in order to make the most informative experiments.
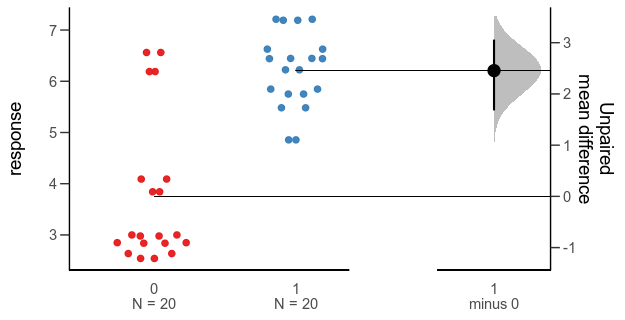
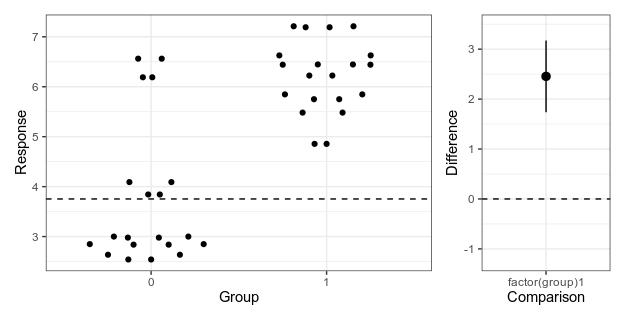
The empirical part of Betts et al. (2021) consists of a literature survey, where the authors read 268 empirical articles from ecology, evolution and glam journals published 1991-2018 to look whether they explicitly stated hypotheses (that is, proposed explanations or causes, regardless of whether they used the actual word ”hypothesis”), whether these were mechanistic, and whether there were multiple working hypotheses contrasted against each other. They estimated the slope over time, and the association between hypothesis use and journal impact factor and whether the research was funded by a major grant.
The results suggest that papers with explicit hypotheses are in a minority, that there was no significant change over time, and little association with impact factor or grants. The prevalence of mechanistic hypotheses was 26% and of multiple working hypotheses 6.7%. There were no significant time trends in hypothesis use. There was a significant difference in journal impact factor in one of the comparisons, where papers with mechanistic hypotheses were published in journals with 0.3 higher impact factor on average. There was no association with grants.
The authors go on to discuss how strong inference is still useful both to the scientific community and to individual researchers, arguing that they might not get more grants or fancier papers, but they will feel better and their research will be of higher quality. How to interpret the lack of clear increase or decrease over time depends on one’s level of optimism I guess. An optimistic take could be that the authors’ fear that machine learning and large datasets are turning researchers away from explanation seem not to be a major concern. A pessimistic take could be, like the they suggest in the Discussion, that decades of admonitions to do hypothetico–deductive science have not had much effect.
Thinking about causality is a good thing
I wholeheartedly agree with the authors that thinking about explanations, causality and mechanism is a useful thing to do, and probably something we ought to do more. It is probably useful to spend more time than we do (for me, to spend more time than I do) thinking about how theories map to testable hypotheses, how those hypotheses map to quantities that can be estimated, and how well the methods and data at hand manage to perform that estimation. Some of my best lessons from science over the last years have come from that sort of thing.
I also agree with them that causality is often what scientists are after — even in many cases where we think that the goal is prediction, the most trustworthy explanation for any ability of a prediction model to generalise is going to be an explanation in terms of mechanisms. They don’t go into this too much, but the caption of Figure 1 gives an example of how even when we are interested in prediction, explanations can be handy.
To take an example from my field: genomic prediction, when we fit statistical models to DNA data to predict trait values for breeding, seems like a pure prediction problem. And animal breeders are pragmatic enough to use anything that worked; if tea leaves worked well for breeding value prediction, they would use them (I am sure I have heard or read some animal breeding researcher make that joke, but I can’t find the source now). But why don’t tea leaves work, while single nucleotide markers spaced somewhat evenly across the genome do? Because we have a fairly well established theory for how genetic variants cause trait variation between individuals in a fairly predictable way. That doesn’t automatically mean that the statistical associations and predictions will transfer between situations — in fact they don’t. But there is theory that helps explain why genomic predictions generalise more or less well.
I also like that they, when they define what a hypothesis is (a proposal of a mechanism or cause of a phenomenon), make very clear that statistical hypotheses and null hypotheses don’t count as scientific hypotheses. There is more to explore here about the relationship between statistical inference and scientific hypotheses, and about the rhetorical move to declare something the null or default model, but that is for another day.
If scientists don’t use strong inference, maybe the problem isn’t with the scientists
Given the mostly negative results, the discussion starts as follows:
Overall, the prevalence of hypothesis use in the ecological and evolutionary literature is strikingly low and has been so for the past 25 years despite repeated calls to reverse this pattern […]. Why is this the case?
They don’t really have an answer to this question. They consider whether most work is descriptive fact finding, or purely about making prediction models, but argue that it is unlikely that 75% of ecology and evolution research is like that — and I agree. They consider a lack of individual incentives for formulating hypotheses, and that might be true; there was no striking association between hypotheses, grants or glamorous publications (unless you consider 0.3 journal impact factor units a compelling individual-level incentive). They suggest that there are costs to hypothesising — it ”an feel like a daunting hurdle”. However, they do not consider the option that their proposed model of science isn’t actually a useful method.
To think about that, we should discuss some of the criticisms of strong inference.
O’Donohue & Buchanan (2001) criticise the strong inference model by arguing that there are problems with each step of the method, and that the history of science anecdotes that Platt use to illustrate it actually show little evidence of being based on strong inference.
Specifically, Platt’s first step, devising alternative hypotheses, is problematic both because one might lack the background knowledge to devise many alternative hypotheses, and that there is no sure way to enumerate the plausible alternative hypotheses.
The second step, devicing crucial experiments, is problematic because of the Duhem–Quine problem, namely that experiments are never conclusive; even when the data are inconsistent with a hypothesis, we do not know whether the problem is with the hypothesis or with any number of, sometimes implicit, auxiliary assumptions. (By the way, I love that Betts et al. cite two ecologists called Quinn and Dunham (1983) who wrote about problems with conclusively testing hypotheses in ecology and evolution. I wish they got together to write it just because the names are so perfect for the topic.)
The third step, conducting crucial experiments, is problematic because experimental results may not cleanly separate hypotheses. Then again, would Platt not just reply that one ought to devise a better experiment then? This objection seems weak. Science is hard and it seems perfectly possible that there are lots of plausible alternative hypotheses that can’t be told apart, at least with data that can be realistically gathered.
Finally, O’Donohue & Buchanan (2001) go through some of Platt’s examples given of supposed strong inference, and suggest that Platt did not represent them accurately. And Platt’s paper really reads as a series of hero-worshipping anecdotes about great scientists, who were very successful and therefore must have employed strong inference. It is not convincing history of science.
Bett et al. (2021) instead give two examples of science that they suppose could have been helped by strong inference. The first example is Lamarck who is supposed to have been able to possibly come up with evolution by natural selection if he had entertained multiple working hypotheses. The second is psychologist Amy Cuddy’s power pose work which supposedly could have been more reproducible had it considered more causal mechanisms. They give no analysis of Lamarck’s scientific method or argument for how strong inference might have helped him. The evidence that strong inference could have helped Amy Cuddy is that she said in an interview that she should have considered the psychological mechanisms behind power posing more.
The claim, inherited from Platt, that multiple working hypotheses reduce confirmation bias really cries out for evidence. As far as I can tell, neither Platt nor Betts et al. provide any, beyond the intuition that you get less attached to one hypothesis if you entertain more than one. That doesn’t seem unreasonable to me, but it just shoves the problem to the next step. Now I have several plausible hypotheses, and I need to decide on one of them, that will advance my decision tree of experiments to the next branching point and provide the headline result for my next paper. That choice seem to me to be just as ripe for confirmation bias and perverse incentives than the choice to call the result for or against a particular hypothesis. In cases where there are only two hypotheses that are taken to be mutually exclusive, the distinction seems only rhetorical.
How Betts et al. (2021) themselves use hypotheses
Let us look at how Betts et al. (2021) themselves use hypotheses and whether they successfully use strong inference for the empirical part of the paper.
That the abstract states two hypotheses — that the number of papers with explicit hypotheses could have decreased because of a perceived rise in descriptive big data research; that explicit hypotheses could have increased because of hypotheses being promoted by journals and funders — none of which turn out to be consistent with the data, which shows a steady low prevalence of explicitly stated hypotheses.
One should note that in no way are these two mechanistic accounts mutually exclusive. If the slope of the line had been positive, that would have no logical force to compel us to believe that the rise of machine learning in research did not lead some researchers to abandon hypothesis-driven research — at most, we could conclude that the quantitative effect of accounts that promote and discourage explicit hypotheses balance towards the former.
Thus, we see two of the objections to Platt’s strong inference paradigm in action: the set of alternative hypotheses is by no means covering the whole range of possibilities, and the study in question is not a conclusive test that allows us to exclude any of them.
In the second set of analyses, measuring whether explicitly stating a hypothesis was associated with journal ranking, citations, or funding, the authors predict that hypotheses ought to be associated with these things if they confer academic success. This conform to their ”if–then” pattern for a research hypothesis, so presumably it is a hypothesis. In this case, there is no alternative hypothesis. This illustrates a third problem with Platt’s strong inference, namely that it is seldom actually applied in real research, even by its proponents, presumably because it is difficult to do so.
If we look at these two sets of analyses (considering change over time in explicit hypothesis use and association between hypothesis use and individual-researcher incentives) and the main message of the paper, which is that strong inference is useful and needs to be encouraged, there is a disconnect. The two sets of hypotheses, whether they are examples of strong inference or not, do not in any way test the theory that strong inference is a useful scientific method, or the normative claim that it therefore should be incentivised — rather, they illustrate them. We can ask Platt’s diagnostic question from the 1964 paper about the idea that strong inference is a method that needs to be encouraged — what would disprove this view? Some kind of data, surely, but nothing that was analysed in this paper.
I hypothesise that this is common in scientific papers. A lot of the reasoning goes on at a higher level than the hypothesis — theories, frameworks, normative stances — and the whether individual hypotheses stand and fall have little bearing on these larger structures. This is not necessarily bad or unscientific, even if it does not conform to Platt’s strong inference.
Method angst
Finally, the paper starts out with a strange anecdote: the claim that there is in the beginning of most scientists’ careers a period of ”hypothesis angst” where the student questions the hypothetico–deductive method. This is stated without evidence, and without following through on the cliff-hanger by explaining how the angst resolves. How are early career scientists convinced to come back into the fold? The anecdote becomes even stranger once you realise that, according to their data, explicit hypothesis use isn’t very common. If most research don’t use explicit hypotheses, it seems more likely that students, who have just sat through courses on scientific method, would feel cognitive dissonance, annoyance or angst over the fact that researcher around them don’t state explicit hypotheses or follow the simple schema of hypothetico–deductivism.
Literature
Betts, MG, Hadley, AS, Frey, DW, et al. When are hypotheses useful in ecology and evolution?. Ecol Evol. 2021; 00: 1-15.
O’Donohue, W., & Buchanan, J. A. (2001). The weaknesses of strong inference. Behavior and philosophy, 1-20.
Platt, JR. (1964) Strong Inference: Certain systematic methods of scientific thinking may produce much more rapid progress than others. Science 146 (3642), 347-353.