(This post was originally written on 2013-03-23. Since then, it has persistently remained one of my most visited posts, and I’ve decided to revisit and update it. I may do the same to some other old R-related posts that people still arrive on through search engines. There was also this follow-up, which I’ve now incorporated here.)
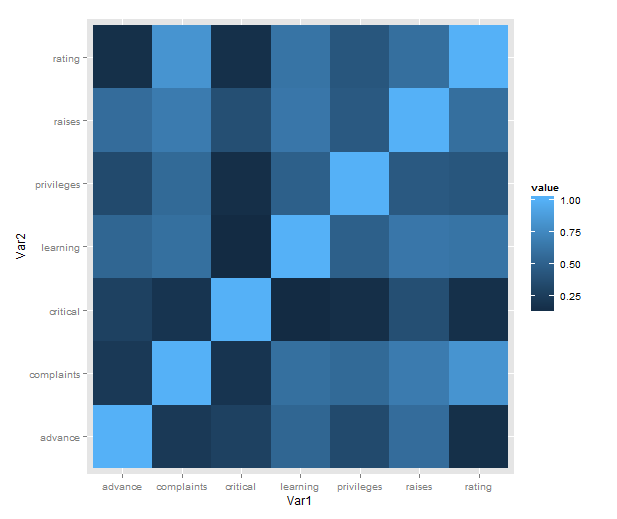
Just a short post to celebrate when I learned how incredibly easy it is to make a heatmap of correlations with ggplot2 (with some appropriate data preparation, of course). Here is a minimal example using the reshape2 package for preparation and the built-in attitude dataset:
library(ggplot2)
library(reshape2)
qplot(x = Var1, y = Var2,
data = melt(cor(attitude)),
fill = value,
geom = "tile")
What is going on in that short passage?
- cor makes a correlation matrix with all the pairwise correlations between variables (twice; plus a diagonal of ones).
- melt takes the matrix and creates a data frame in long form, each row consisting of id variables Var1 and Var2 and a single value.
- We then plot with the tile geometry, mapping the indicator variables to rows and columns, and value (i.e. correlations) to the fill colour.
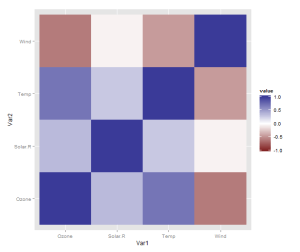
However, there is one more thing that is really need, even if just for the first quick plot one makes for oneself: a better scale. The default scale is not the best for correlations, which range from -1 to 1, because it’s hard to tell where zero is. Let’s use the airquality dataset for illustration as it actually has some negative correlations. In ggplot2, a scale that has a midpoint and a different colour in each direction is called scale_colour_gradient2, and we just need to add it. I also set the limits to -1 and 1, which doesn’t change the colour but fills out the legend for completeness. Done!
data <- airquality[,1:4]
qplot(x = Var1, y = Var2,
data = melt(cor(data, use = "p")),
fill = value,
geom = "tile") +
scale_fill_gradient2(limits = c(-1, 1))
Finally, if you’re anything like me, you may be phasing out reshape2 in favour of tidyr. If so, you’ll need another function call to turn the matrix into a data frame, like so:
library(tidyr)
correlations <- data.frame(cor(data, use = "p"))
correlations$Var1 <- rownames(correlations)
melted <- gather(correlations, "value", "Var2", -Var1)
qplot(x = Var1, y = Var2,
data = melted,
fill = value,
geom = "tile") +
scale_fill_gradient2(limits = c(-1, 1))
The data preparation is no longer a oneliner, but, honestly, it probably shouldn’t be.
…
Okay, you won’t stop reading until we’ve made a solution with pipes? Sure, we can do that! It will be pretty gratuitous and messy, though. From the top!
library(magrittr)
airquality %>%
'['(1:4) %>%
data.frame %>%
transform(Var1 = rownames(.)) %>%
gather("Var2", "value", -Var1) %>%
ggplot() +
geom_tile(aes(x = Var1,
y = Var2,
fill = value)) +
scale_fill_gradient2(limits = c(-1, 1))